Biography
I build AI systems that model how cells respond to drugs and perturbations — bridging deep generative models, single-cell biology, and production ML to accelerate therapeutic discovery.
At AI VIVO I develop models for molecular generation (SMILES, ADMET), structure-based design (AutoDock Vina, Boltz-2), virtual cells and perturbation prediction and generation, and multimodal pipelines at scale. As a Data Scientist at the Wellcome Sanger Institute (Lotfollahi Lab), I co–first authored CellDISECT (bioRxiv 2025; under review at Nature Methods), co-authored SP-FM (arXiv 2026), fine-tune single-cell foundation models for perturbation prediction, and help maintain CPA for the community.
- Deep Generative Models
- Causal Representation Learning
- Flow Matching
- Single-Cell Perturbation
- Virtual Cells
- Drug Discovery
Applied Computer Science & Artificial Intelligence, 2026
Sapienza University of Rome
BSc in Computer Science, 2023
Amirkabir University of Technology
Current Work
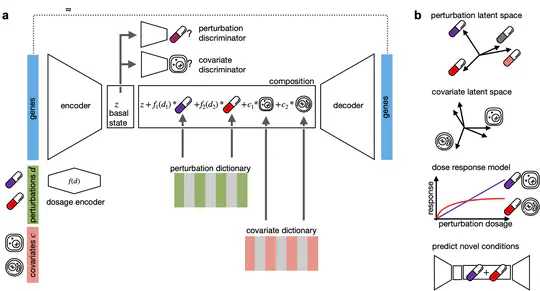
- CellDISECT (bioRxiv, 2025; under review at Nature Methods) / Code: Causal VAE for covariate disentanglement, counterfactual perturbation prediction, and cell-type discovery from large-scale scRNA-seq.
- SP-FM (arXiv:2601.11827, 2026): Shortest-path flow matching with mixture-conditioned bases for OOD generalization to unseen biological conditions.
- Why Perturbation Prediction Needs Much Better Metrics (Medium, Mar 2026): Essay on evaluation metrics, mean-predictor baselines, and benchmarking in computational biology.
- AI VIVO: Machine learning for de novo molecular generation, genetic and chemical perturbation prediction (virtual cells), and target discovery.
Work Experience
- Develop deep learning and generative models for drug discovery using transformer and flow matching architectures
- Deploy and scale ML pipelines on GCP using PyTorch Lightning and Docker
- Design multi-modal ML pipelines integrating molecular structure and biological assay data
- Maintain scalable pipelines using PyTorch, Lightning, RDKit, and HuggingFace
- Experience with computational chemistry tools like BioSolveIt, AutoDock Vina, and Boltz-2
- Lotfollahi Lab: co–first author of CellDISECT — causal VAE for disentangled single-cell representations and in silico perturbation prediction (2M+ cells, 100+ counterfactual conditions); manuscript under review at Nature Methods
- Fine-tuning of ~300M-parameter single-cell transformer foundation models (e.g. scFoundation, Geneformer, scGPT) with LoRA and prompt tuning on 1500+ perturbation conditions
- Lead maintainer for CPA (Compositional Perturbation Autoencoder), including high-volume code review and releases for the single-cell community
- Collaboration across a large PhD/postdoc team on preprints, open-source tools, and PyTorch-based single-cell genomics methods
Responsibilities include:
- Delivered >95% accuracy solutions for tasks with limited data (15 images per class)
- Spearheaded development on 5 diverse projects meeting client requirements
- Led individual projects, enhancing development pipelines
Featured Publications
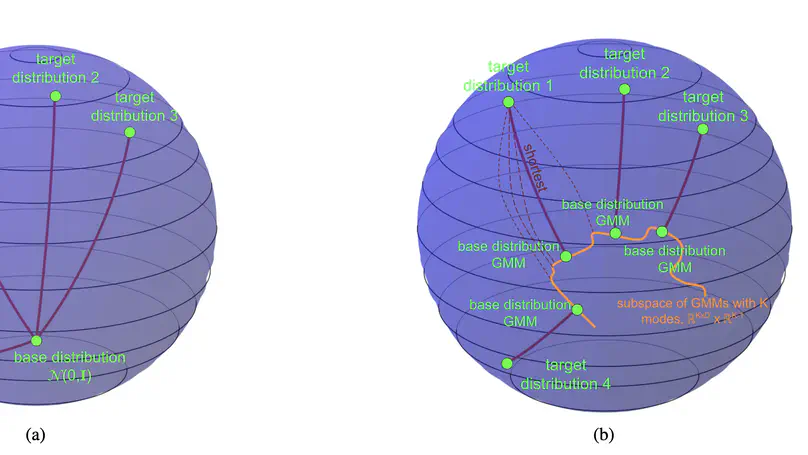
SP-FM introduces a shortest-path flow-matching framework with mixture-conditioned bases to improve out-of-distribution generalization in conditional generative modeling, enabling better extrapolation to unseen conditions.
Projects
Recent Publications
Recent & Upcoming Talks
Teaching Experience
- Machine Learning for Bioinformatics (Graduate Course) | Spring 2023
- Prepared teaching material on CNNs & AutoEncoders, designed assignments, and coordinated class contests.
- Introduction to Machine Learning | Fall 2022
- Designed and graded assignments for a class of 150 students, conducted a workshop on Variational AutoEncoders.
- Introduction to Image Processing and Neural Networks | Fall 2022
- Conducted workshops and lectures on OpenCV and Deep Learning for a class of 80 students.
- Advanced Programming with C++ | Spring 2022
- Designed assignments and projects for a class of 90 students, evaluated student submissions.
Accomplishments
- Neural Networks and Deep Learning
- Improving Deep Neural Networks: Hyperparameter Tuning, Regularization and Optimization
- Structuring Machine Learning Projects
- Convolutional Neural Networks
- Sequence Models
Contact
- arianamaani@gmail.com
- Rome, Italy
- DM Me
- Contact Me